|
Authors: Paula Brígido, Roberto de la Fuente and Javier Del Río Assisted reproduction technology (ART) can help fertile couples to achieve successful pregnancies. Sometimes, reproductive desires of these couples are affected by the presence of a genetic disease in either partner. In such cases, couples are at a reproductive risk and find themselves in the need of assistance that only ART can provide. Preimplantation genetic diagnosis (PGD) provides an alternative to prenatal diagnosis to detect the specific genetic condition or disease they suffer from, and allows them to avoid passing it on their offspring (2). It requires the analyses of the embryos generated by ART in the in vitro fertilization (IVF) laboratory, by means of accurate and sensitive methodologies such as embryo biopsy, genetics, single cell genomics and, of course, background on prenatal diagnosis and counselling from experts. Clinical application of PGD dates back to the late 60’s, when blastocysts of research animals could be sexed (3) (note that this was already possible ten years before Louis Brown, the first IVF baby, was born in the UK in 1978). At the beginning of the 90’s, early human embryos were sexed before implantation and the first genetic analyses were performed to avoid children inheriting Mendelian diseases. By the end of the century, other nowadays considered basic genetic methodologies were routinely used for preimplantation diagnosis and PGD was applied as a normal procedure to guarantee healthy babies (4). In the present post we aim to give an account of the importance of PGD and the current view of the main clinical approaches for its application. WHEN IS PGD INDICATED? Indications for PGD are multiple and emerge from different motivations. Firstly, the patient may have suffered from a number of terminations due to the embryo having inherited the genetic condition. It could also be motivated by the parents already having a child with a severe genetic disease. In this case they might be willing to avoid passing it on the next one or even looking for a suitable treatment, if possible. However, one of the parents (or both) may be worried about their family history, being aware of the presence of a specific genetic condition, regardless of the type of inheritance. If the parents are carriers of any genetic disease, either an autosomal-dominant disorder like Huntington disease or an autosomal-recessive one like cystic fibrosis, they are at reproductive risk because the resulting embryo may be affected (the probability depending on the specific disorder itself and the way it is inherited) (see [2] for details on inherited conditions). But there are even cases in which motivation is not based on biological but in ethical or religious reasons. Certain families might have serious concerns about going on for abortion of an affected embryo. In such cases, application of PGD may circumvent this kind of ethical conflicts. Applying PGD Broadly speaking, steps for PGD are as follows (2):
PGD vs. PGS Preimplantation genetic screening (PGS) is the general term for a compound of approaches that aim to evaluate the genetic content of the cell, in contrast to genetic tests whose goals are to determine whether an embryo is affected by a specific genetic condition (PGD). Originally termed PGD-AS (preimplantation genetic diagnosis for aneuploidy screening), PGS was developed to confirm the ploidy status of the embryo, searching for possible aneuploidies. Available data suggest that most of miscarriages occurred during the first trimester are a consequence of some sort of aneuploidies (5), and that mainly selected chromosomes were involved in these structural abnormalities (6). Thus, the main approach developed for PGS was the fluorescence in situ hybridization (FISH) for such chromosomes. Types of approaches for PGD in the laboratory Current technical methodologies for preimplantational genetic analyses mainly lie in one of the following:
WHEN TO PERFORM BIOPSY Typical biopsies for PGD (and PGS) are as described as follows:
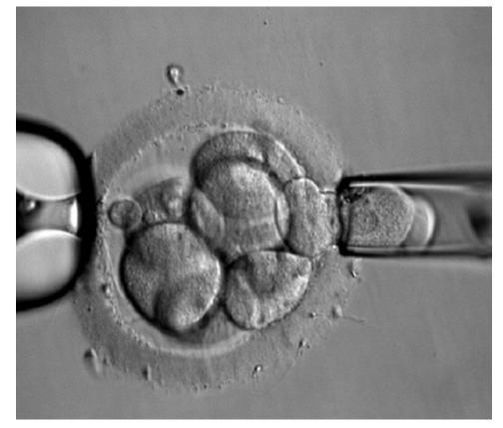
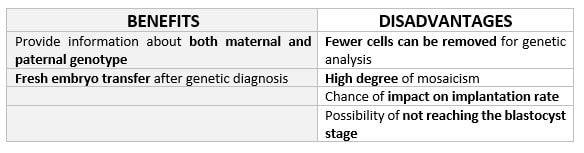
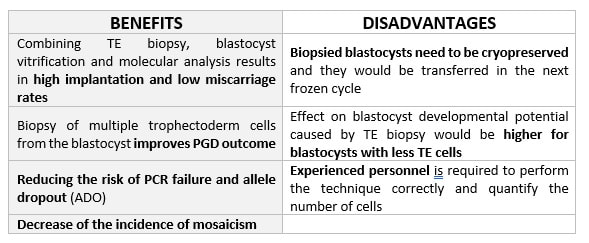
DAY 3. CLEAVAGE STAGE BIOPSY There is a controversy regarding utility of this type of biopsy. In the cleavage stage biopsy, embryos are biopsied at day 3 when individual cells can be differentiated. This technique entails aspiration of one to two blastomeres to obtain the embryonic genetic material for PGD analysis (13). Following genetic diagnosis, embryo transfer may be performed on blastocyst stage. Embryos are usually selected for biopsy based on morphological criteria. Unfortunately, these do not predict the development potential of the embryo, and so it could fail to progress until blastocyst stage. This would compromise the advantages of using the day-3 approach (14). On the other hand, performing biopsy on the cleavage stage allows embryos to be cultured in vitro until they reach the blastocyst. This means they can be fresh transferred (15), whereas embryos biopsied on day 5 must be vitrified and transferred in a subsequent cycle. How many cells should be removed? The number of cells to be removed in the biopsy is still a controversial issue. Aspirating one cell reduces the cellular mass extracted but it can imply the presence of mosaicism. Conversely, aspirating two cells can reduce the risk of mosaicism, but removing such cellular mass could have consequences on the implantation rate (14). Reported data have shown a dramatic reduction of 39% in the implantation rate in cleavage stage biopsy (16). The authors related it with proportion of the embryo total cellular removed. Whereas around five cells pulled out of the embryo in the trophectoderm biopsy represent 2-3% of the total cell content (expanded blastocyst has 200-220 cells approximately), extraction of a single cell from an eight cell embryo supposes 13% of the total content (16). What do experts say? Cleavage stage biopsy produces different opinions among embryologists because of the presence of mosaicism and the possibility of self-correction of aneuploidies from cleavage to blastocyst stage (17). On the contrary, studies using array-comparative genomic hybridization (array-CGH) technology to analyse genetic abnormalities in day-3 blastomeres and confirming it in trophectoderm biopsy showed concordance between day 3 diagnosis and day 5 reanalysis; Treff and coauthors showed more reliable results for SNP-microarray (96% vs. 83%) and also a lower mosaicism degree (31%) for SNP-microarray samples in a study comparing array technology versus FISH technique (18). These data would support the suggestion of some authors, who proposed that the incidence of mosaicism may have been overestimated in previous studies due to technical inconsistency of the FISH technique (17, 18, 19). At present, this matter remains controversial. Regarding pregnancy rates, in both types of biopsies higher pregnancy rates are obtained comparing with the control group, in which no biopsy was performed (14, 19). To sum up: DAY 5. TROPHECTODERM BIOPSY The blastocyst stage is currently supposed to be an optimal time to perform biopsies for PGD/PGS. The combination of improved blastocyst culture, trophectoderm (TE) biopsy, refined cryopreservation techniques, and molecular assays, such as array comparative genomic hybridization that allows for 24-chromosome screening, have led to a renaissance of PGS. TE biopsy will not detect every circumstance in which the embryo is at risk of aneuploidy, but it will detect mosaicism more reliably than cleavage-stage biopsy (which cannot be relied on at all for this purpose) (20, 21). Moreover, when diagnosing monogenic disorders in single blastomere cells using PCR-based protocols, there is a high risk of PCR failure due to either no amplification (allele dropout) or preferential amplification of one of the alleles, potentially resulting in a reduced number of unaffected embryos available for transfer. Increasing the amount of starting DNA template should in principle increase the sensitivity and reliability of genetic diagnosis. Therefore, the biopsy of multiple trophectoderm cells from the blastocyst rather than a single cell from cleavage stage embryos should potentially lead to improved PGD outcome for patients (14). How many cells should be removed? Research to determine the appropriate number of biopsied TE cells in blastocyst biopsies are limited. The exact number of biopsied TE cells is hard to count visually because cells are small and usually remain as a clump. In most studies using comparative genome hybridization or single-nucleotide polymorphism array technology for genetic testing, biopsied TE cells were used for genome amplification and their number was impossible to know. Moreover, some studies showed that removing four to five cells leads to better results. Therefore, the biopsied cell number should be higher in the blastocysts with better TE quality than those with worse characteristics (22, 23). Can biopsies affect blastocyst development and its implantation? Whereas it remains possible that biopsy of cleavage-stage embryos can critically arrest further development through reduction of cell mass, the low miscarriage rates and high term birth rates in the present series, as well as data presently under analysis, suggest that this is not the case for TE biopsy. It can be speculated that the damage to blastocyst development potential caused by TE biopsy would be less for blastocysts with a greater number of TE cells (21, 22). Some experts assured that TE biopsy at the blastocyst stage had no meaningful impact on the developmental competence of the embryo as measured by implantation and delivery rates. This contrasts with the information above-mentioned on the significant reduction in the probability for an embryo to implant and progress up to delivery (16). When combined with TE biopsy and blastocyst vitrification, SNP microarray has resulted in high implantation and low miscarriage rates for some IVF patients (15, 16, 24). Are there any limitations? Owing to the limitations of genetic analysis, most of the biopsied blastocysts need to be cryopreserved by vitrification, and blastocysts with normal results would be transferred in the next frozen cycle. In addition, biopsy of numerous cells from blastocysts with grade B or C may cause damage to the embryo, leading to either its arrest or implantation failure. However, 1-5 cells may be the appropriate biopsied TE cell number to maintain the implantation potential (15, 22). Also, the personnel experience of different embryologists is an influencing factor in this technique. The number of biopsied cells in the blastocyst biopsy is hard to quantify and largely dependent on the experience of embryologist (22). To sum up: WHAT CAN WE CONCLUDE? The availability of new embryology and molecular techniques allow preimplantation genetic diagnosis laboratories to offer patients at genetic risk the transfer of developmentally competent embryos, unaffected by genetic disease. Cleavage stage biopsy allows for fresh embryo transfer after genetic diagnosis. However, there are reports of high levels of mosaicism when the biopsy is performed on day 3. Trophectoderm biopsy, in turn, provides sufficient material for an effective and more reliable diagnosis in embryos compared to those on cleavage stage. Moreover, it seems that it does not compromise embryo implantation and pregnancy rates in PGD cycles. The drawback for this option is the usual need for cryopreservation and transfer in a different cycle. The offer of PGD in fertility centres has increased over the last decade, primarily due to the progress on the application of diagnostic methods. The choice for either development stage relates to successful outcomes in the clinic, which mainly depend on technical challenges and timing of the developing embryo. For the embryologists, both day-3 and day-5 approaches are supported by evidence, but it will be essential to consider every single aspect of them to evaluate the best option for the laboratory. REFERENCES:
4 Comments
|
Entries
March 2019
Categories
All
 2016-2019. All Rights Reserved by Embryologist Media. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License . |
Embryologist Media