|
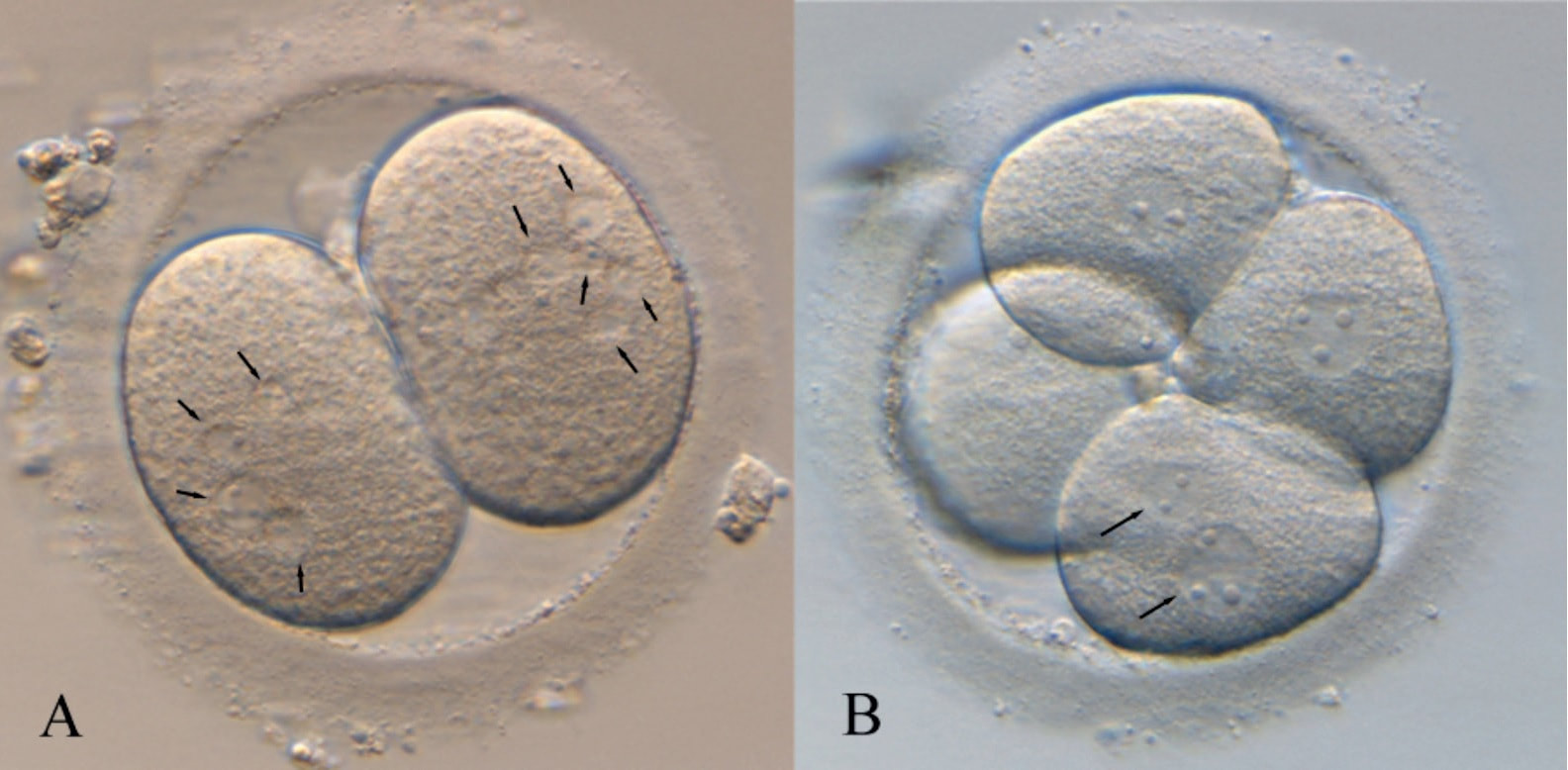
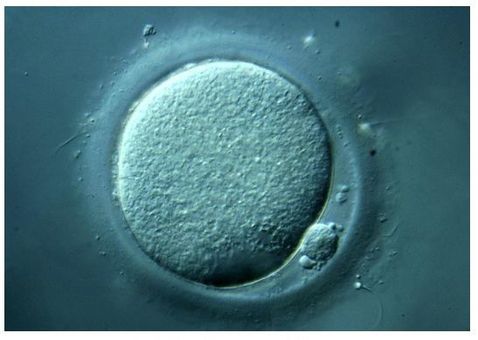
Authors: Iris Martínez Rodero and Raquel Pillado González "Multinucleated blastomeres (MNBs) present in embryos are morphological abnormalities of unclear origin, which have been extensively correlated with chromosomal defects, lower blastocyst formation and implantation rates". INTRODUCTION Selection of high-quality embryos is an important factor for the successful outcome of assisted reproduction technologies (ART). Nowadays, criteria for selection are mainly based on morphological features such as embryo fragmentation, cell number, blastomeres uniformity, etc. (2). The parameters studied so far have been demonstrated to be useful indicators of embryo quality. Their evaluation is performed through non-invasive light microscopy-based analyses, usually carried out once a day at specific time points. This approach intends to minimise the events of taking the embryos out of the incubators and exposing them to undesired harmful conditions. The presence of multinucleated blastomeres (MNBs) can be regarded as one of those indicators, and even though previous studies had already connected it to DNA abnormalities and low pregnancy rates as early as in the 90s (3,4,5) the origin of this phenomenon still remains unclear. Several possible factors seem to influence its genesis, but the specific cause for its occurrence is yet to be determined (6). Since the introduction of time-lapse imaging and monitoring technology, IVF laboratories have been able to carry out more exhaustive and continuous observations on embryo development, keeping risks at a minimum (7,8). By identifying the precise timing of specific key events of blastomere cell cycle interludes and of the embryo´s overall growth it is possible to assess its quality (7). Furthermore, time-lapse imaging and monitoring systems have facilitated the study of multinucleation (MN) in relation to its incidence in time and in the population, as well as its correlation with other morphological features and clinical variables (6,7,9). In the IVF context multinucleation is defined as the presence of two or more nuclei in one or more blastomeres. Multiples studies even differentiate between binucleated and multi-/micro-nucleated (three or more nuclei) blastomeres (Fig 1) (1,4,7). Reports on this phenomenon range from 17% to 69% of the total of cultured embryos, depending on the groups used within assays and authors (10). Factors that may influence MNB appearance are numerous and have been repeatedly studied (6,7,9,11). To date, several explanations have been proposed for MNBs: dysfunction of the mitotic spindle or the occurrence of karyokinesis without cytokinesis (12); DNA breaks or imperfect mitosis (13); nuclear membrane alterations (14) and even other factors not directly responsible for MNBs and yet linked to its presence (1). FACTORS THAT MAY RELATE TO THE APPEARANCE OF MULTINUCLEATION
Results from various studies have shown a difference in the percentage of multinucleated embryos between groups that had been fertilised by traditional IVF vs ICSI. Van Royen and colleagues showed 32.7% of MN embryo in the IVF group, compared to 34.5% in the ICSI group (6). Accordingly, Walmsley et al. 2003 reported 17,2% vs 18,3% of MN in embryos derived from IVF and ICSI, respectively (11).
Apparently, the type of infertility factor seems not to affect MN rates. Some studies have reported no significant differences in the percentages of MN embryos between cases of female factor-only infertility and male factor-only infertility (32.7% and 34.7%, respectively). In addition, differences were not found between these cases and those with both partners affected by some sort of infertility, either (6). However, different studies support the relevance of oocyte culture, specially regarding certain processes that occur naturally in in vivo conditions and that are essential for the proper embryo development. Data show that both oocytes subjected to negative conditions when cultured in vivo (like stern hypoxia, for instance) and oocytes cultured in vitro derive in a higher percentage of MN embryos (15,16). Regarding male infertility, results are more controversial; whereas some studies reflect a higher MN rate for cases in which male factor is especially severe (normally derived for ICSI rather than IVF) (11), other authors show no significant differences (6).
Records exhibit that cycles with an accelerated ovulation induction response present increased MN rates (6,17). Furthermore, several studies have reported that embryos derived from patients from whom ten or more oocytes had been collected presented a significantly higher MN rate than embryos from groups of nine or fewer oocytes (6,8,18). This is in accordance with the fact that patients who need high doses of FSH present higher MN rates (6). Very short cycles and cases where high doses of GnRH are needed trigger the development of high numbers of premature follicles that produce oocytes, which despite being able to reach metaphase II and become fertilized fail to go through proper nuclear cleavage (6,18). Lastly, even though differences in GnRH doses have been associated to significant differences in the incidence of MN embryos, similar results have not been observed when using different hormones (like rFSH, r-hFSH, purified urinary FSH or urinary gonadotropin, for instance) (6).
Multiple studies have been conducted on patients ranging from 25 to 45 years old. Several authors divided data in 5-year interval groups in order to verify whether patient age correlates with MN. However, the only significant difference was found when comparing women of +40 with younger ones of -35, presenting higher degree of MN in the first group (6,9).
Chromosome polymorphisms consist in heterochromatin variability. These are usually located in the long arms of chromosomes 1, 9 and 16, and the short arms of chromosomes from the groups D and G (13, 15 and 21, 22 and Y) (19). Even though such polymorphisms are generally regarded as normal within karyotypes (20), studies indicate that some of them might be associated with certain clinical problems such as abnormal spermatogenesis (21), infertility (22,23), recurrent miscarriages (24,25) and higher rate of chromosome abnormalities among blastomeres at the cleavage stage (26,27). Sun and collaborators hypothesized that couples with chromosome polymorphisms might experience a higher rate of embryo multinucleation (19). Nevertheless, the authors found no association between chromosome polymorphisms and MN embryo formation in couples undergoing IVF (19).
- Cellular fragmentation Although MN may appear regardless of the cellular fragmentation levels, several papers support the correlation between these two features (6,28,29). In particular, Van Royen et al. divided the level of fragmentation in three categories: F1 (≤10%), F2 (10-20%) and F3 (20-30%); this study presented evidence for higher MN in F2 and F3 when compared to F1, but similar to each other (6). - Cleavage rate When 3-cell and 5‐cell day-2 embryos were observed under the microscope, both types exhibited significantly higher multinucleation (28.2% -50%) than regular ones with the ideal 4-cell cleavage pattern (with only 16.8% MN). Similarly, day-3 embryos with the typical 8‐cell stage showed significantly lower multinucleation (15.5%) than 7‐cell and 9‐cell embryos (6). As it has been mentioned, ideal 4-cell and 8-cell stages show similar MN percentages. However, application of time-lapse imaging has revealed a significant decrease in MN from the 2-cell to the 4-cell stage (from 43.2% to 15.0%). The analysis of MN in 2-cell embryos indicated that, after cleavage, the majority (52%) of 2-cell MN embryos became mononucleated, whereas only a lower percentage (34%) showed MNBs, and about 14% were of poor quality (with only one or no visible nucleus at all) (9). This decrease in the MN rate suggests that 2-cell MN embryos are able to self-correct their nuclear abnormalities. But this repair mechanism has been observed in both euploid and aneuploid embryos, therefore it cannot be used as an indicator of chromosomal normality during embryo selection (8,9,30). An extended duration of both 2-cell and 4-cell stages has been proposed as a possible indicator of the occurrence of nuclear self-correction (9). INCIDENCE OF MULTINUCLEATION IN CLINICAL IVF As previously exposed, it is through time-lapse imaging that a far higher percentage of multinucleation (25%) has been detected compared to static observations on day 2 at 42 hours post-insemination (hpi) (<5%) (7). These observations have demonstrated that multinucleation is a frequent event that, according to Yilmaz et al., is present in at least one embryo in 41.3% of IVF cycles (31). Data provided by Desai and colleagues reported that approximately 56% of binucleated embryos and 48% of those with three or more nuclei went on to form blastocysts that met the appropriate criteria for vitrification (7). In addition, data from different studies point to binucleation being more frequent than blastomeres with 3 or more nuclei (7,31,32). At the same time, multinucleation has provided an additional criterion for embryo selection, since it is mainly observed in those of poor quality and is associated with direct and/or reverse cleavage (7). It has been observed that, out of all embryos found showing direct and/or reverse cleavage, at least one fourth were also multinucleated (7). By using time-lapse, multinucleation has been repeatedly observed to be a reversible event in a high proportion of embryos (7,32). Multinucleation reversibility has been reported to be as high as 73.4% (32); this has been calculated as the proportion of embryos in which multinucleation was detected at 2-cell stage, but not visible at 4-cell stage (likely due to self-correction mechanisms, as above-mentioned). In fact, Aguilar and collaborators reported 127 multinucleated embryos at 4-cell stage out of the 479 ones initially observed to present this feature at 2-cell stage. De novo multinucleation at the 4-cell stage in turn was observed in 36 embryos (32). IMPACT OF MULTINUCLEATION ON IVF OUTCOMES Multinucleation has traditionally been related to both low blastocyst formation (33) and implantation rates (5,6,17,28,34), and linked to the likely presence of chromosome abnormalities, which consequently results in embryo arrest (35). Nevertheless and despite all the existing evidences, there is still much controversy regarding multinucleation; reports have been published revealing cases in which fully binucleated 4-cell stage embryos had eventually developed into euploid blastocysts and genetically normal children (31,34).
Although some preimplantation genetic testing (PGT) studies have shown that not all multinucleated embryos are chromosomally abnormal (31,32,36) multinucleation is predominantly associated to chromosomal defects and poor implantation prognosis (3,31,37). Kligman and colleagues published that 74.5% of multinucleated embryos were chromosomally abnormal, compared to 32.3% of non-multinucleated embryos (3). Years later, Ambroggio et al. revealed an increased incidence of aneuploidy of MN 4-cell stage embryos when compared to single-nucleated embryos (85% vs 78%), suggesting that multinucleated embryos should not be recommended for transfer in IVF cycles (37). These results were confirmed when, from 395 MN embryos tested for PGT, Yilmaz et al. reported that 82.5% of MN blastomeres exhibited two nuclei, whereas the remaining blastomeres presented a single or three or more nuclei (31). Noteworthy, binucleated patterns of multinucleation may be less detrimental, since a high percentage of embryos with such feature are euploid, compared to embryos exhibiting three or more nuclei in a single blastomere (38).
Embryo morphokinetics was studied and related to the multinucleation status in a study conducted by Meseguer’s team (32). In the study, 53.4% of a total 1676 embryos included were MN. Based upon the reported data, differences in morphokinetics between multinucleated and non-multinucleated embryos at both 2-cell and 4-cell stages comprise cleavage events involving the completion of the first mitosis and the length of the S-phase. These differences affected the following parameters: t2, t3, second cell cycle (cc2=t3-t2), t4, t6, t7 and t8. These results allowed to conclude that, if multinucleation remains at 4-cell stage, it takes longer for the embryos to complete the next cell cycle (cc3=t5-t3). Should this be true, the restoration system would not be efficient if MNBs were still observed after the 2-cell stage (32). The origin of the multinucleation phenotype has been suggested to be multiple: disruption of intracellular restructuring, remodelling or imprinting in the developing oocyte, or even alterations in DNA replication, cytokinesis or compaction during the first cell cycle (16). If multinucleation appears as a result of defects in cell function, differences in morphokinetics between MN and non-MN embryos during these early stages may be expected (32).
Opinions on the impact of the multinucleation phenotype on implantation rates diverge from each other: On one hand, cell stage for MN appearance has been proposed to exert the highest effect on the implantation rate. Authors supporting this claim are divided into two positions: those who affirm that the presence of MN at the 2-cell stage is actually insignificant in terms of differences on implantation rates, but it is at 4-cell stage when it does have a measurable negative effect (32); and the authors who argue implantation rates to be significantly reduced when MN is already observed at the 2-cell stage (8). On the other hand, the school of Meriano and coauthors affirm that binucleation is less harmful than any other type of multinucleation (16). However, Aguilar and colleagues explained that their differences with Meriano were found on the frequency of image acquisition and the systems used to measure multinucleation (32); whereas the former acquired one picture in seven different focal planes every 20 minutes, the latter recorded images every 2.5 minutes (32). In any case, it has been demonstrated that patterns of multinucleation at 4-cell stage are correlated with low implantation rates, while any of the other cases has been reported to decrease the chances to achieve pregnancy (16,32). CONCLUSIONS Multinucleation is a common and reversible event observed in human IVF embryos, and it is specially frequent as binucleation at two-cell stage. It is associated with chromosomal defects and altered morphokinetic parameters, eventhough binucleation patterns seem to be the less severe. Regarding multinucleation impact on implantation rate, results are controverted. It seems that implantation rates are not affected when multinucleation appears as two nuclei in two-cell stage. Although the presence of multinucleated blastomeres in human embryos has been associated with the above-mentioned undesired characteristics in IVF embryos, the reasons explaining its appearance and occurrence in time and its relationship with patient specifications have not been deeply studied until time-lapse systems became available. Even though different causes have been suspected to lie behind MNB development, none of them have been proved actually represent the main responsible. Nevertheless, a growing number of studies provide data untangling the relationship between MN and assisted reproduction fertilization methods (IVF and ICSI), stimulation cycles, infertility factors, culture conditions and other embryonic morphological characteristics. Even though sometimes results from different studies may seem contradictory, this might be accredited to the differences in sample sizes. All the above said, it seems reasonable to highlight the need for further research on this issue. It would be highly helpful to unveil the actual triggers of multinucleation, to develop optimal ART practices that avoid increasing MN incidence, and to unravel any other correlation with adverse embryo features during development. Deeper knowledge would help improving embryo assessment methods and, consequently, increase the rates of successful ART outcomes. REFERENCES
0 Comments
Authors: Paula Brígido, Roberto de la Fuente and Javier Del Río Assisted reproduction technology (ART) can help fertile couples to achieve successful pregnancies. Sometimes, reproductive desires of these couples are affected by the presence of a genetic disease in either partner. In such cases, couples are at a reproductive risk and find themselves in the need of assistance that only ART can provide. Preimplantation genetic diagnosis (PGD) provides an alternative to prenatal diagnosis to detect the specific genetic condition or disease they suffer from, and allows them to avoid passing it on their offspring (2). It requires the analyses of the embryos generated by ART in the in vitro fertilization (IVF) laboratory, by means of accurate and sensitive methodologies such as embryo biopsy, genetics, single cell genomics and, of course, background on prenatal diagnosis and counselling from experts. Clinical application of PGD dates back to the late 60’s, when blastocysts of research animals could be sexed (3) (note that this was already possible ten years before Louis Brown, the first IVF baby, was born in the UK in 1978). At the beginning of the 90’s, early human embryos were sexed before implantation and the first genetic analyses were performed to avoid children inheriting Mendelian diseases. By the end of the century, other nowadays considered basic genetic methodologies were routinely used for preimplantation diagnosis and PGD was applied as a normal procedure to guarantee healthy babies (4). In the present post we aim to give an account of the importance of PGD and the current view of the main clinical approaches for its application. WHEN IS PGD INDICATED? Indications for PGD are multiple and emerge from different motivations. Firstly, the patient may have suffered from a number of terminations due to the embryo having inherited the genetic condition. It could also be motivated by the parents already having a child with a severe genetic disease. In this case they might be willing to avoid passing it on the next one or even looking for a suitable treatment, if possible. However, one of the parents (or both) may be worried about their family history, being aware of the presence of a specific genetic condition, regardless of the type of inheritance. If the parents are carriers of any genetic disease, either an autosomal-dominant disorder like Huntington disease or an autosomal-recessive one like cystic fibrosis, they are at reproductive risk because the resulting embryo may be affected (the probability depending on the specific disorder itself and the way it is inherited) (see [2] for details on inherited conditions). But there are even cases in which motivation is not based on biological but in ethical or religious reasons. Certain families might have serious concerns about going on for abortion of an affected embryo. In such cases, application of PGD may circumvent this kind of ethical conflicts. Applying PGD Broadly speaking, steps for PGD are as follows (2):
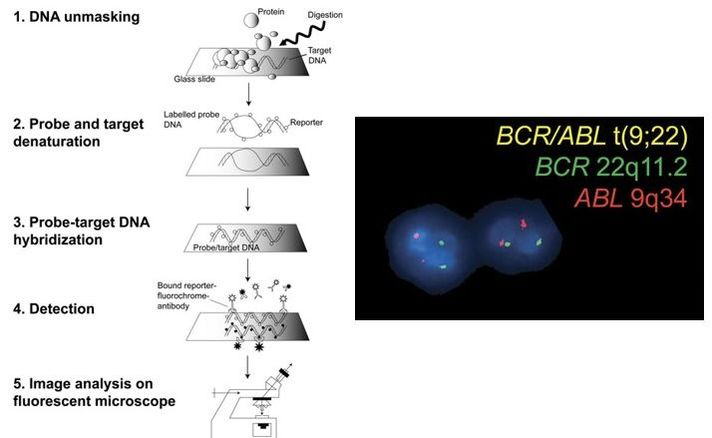
PGD vs. PGS Preimplantation genetic screening (PGS) is the general term for a compound of approaches that aim to evaluate the genetic content of the cell, in contrast to genetic tests whose goals are to determine whether an embryo is affected by a specific genetic condition (PGD). Originally termed PGD-AS (preimplantation genetic diagnosis for aneuploidy screening), PGS was developed to confirm the ploidy status of the embryo, searching for possible aneuploidies. Available data suggest that most of miscarriages occurred during the first trimester are a consequence of some sort of aneuploidies (5), and that mainly selected chromosomes were involved in these structural abnormalities (6). Thus, the main approach developed for PGS was the fluorescence in situ hybridization (FISH) for such chromosomes. Types of approaches for PGD in the laboratory Current technical methodologies for preimplantational genetic analyses mainly lie in one of the following:
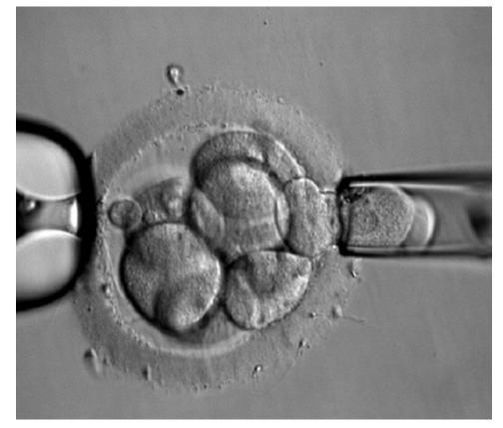
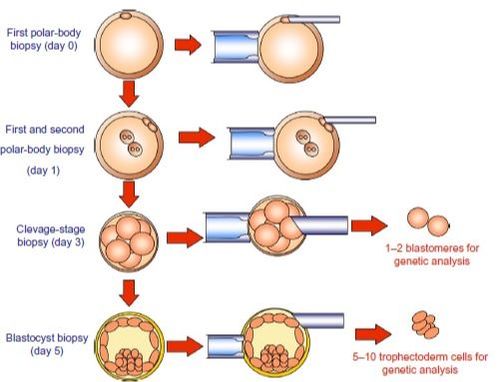
WHEN TO PERFORM BIOPSY Typical biopsies for PGD (and PGS) are as described as follows:
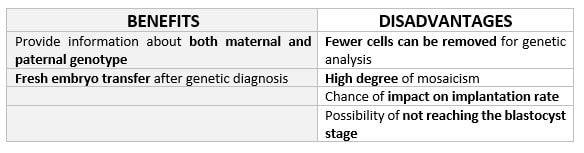
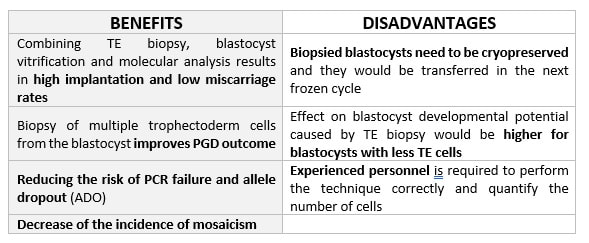
DAY 3. CLEAVAGE STAGE BIOPSY There is a controversy regarding utility of this type of biopsy. In the cleavage stage biopsy, embryos are biopsied at day 3 when individual cells can be differentiated. This technique entails aspiration of one to two blastomeres to obtain the embryonic genetic material for PGD analysis (13). Following genetic diagnosis, embryo transfer may be performed on blastocyst stage. Embryos are usually selected for biopsy based on morphological criteria. Unfortunately, these do not predict the development potential of the embryo, and so it could fail to progress until blastocyst stage. This would compromise the advantages of using the day-3 approach (14). On the other hand, performing biopsy on the cleavage stage allows embryos to be cultured in vitro until they reach the blastocyst. This means they can be fresh transferred (15), whereas embryos biopsied on day 5 must be vitrified and transferred in a subsequent cycle. How many cells should be removed? The number of cells to be removed in the biopsy is still a controversial issue. Aspirating one cell reduces the cellular mass extracted but it can imply the presence of mosaicism. Conversely, aspirating two cells can reduce the risk of mosaicism, but removing such cellular mass could have consequences on the implantation rate (14). Reported data have shown a dramatic reduction of 39% in the implantation rate in cleavage stage biopsy (16). The authors related it with proportion of the embryo total cellular removed. Whereas around five cells pulled out of the embryo in the trophectoderm biopsy represent 2-3% of the total cell content (expanded blastocyst has 200-220 cells approximately), extraction of a single cell from an eight cell embryo supposes 13% of the total content (16). What do experts say? Cleavage stage biopsy produces different opinions among embryologists because of the presence of mosaicism and the possibility of self-correction of aneuploidies from cleavage to blastocyst stage (17). On the contrary, studies using array-comparative genomic hybridization (array-CGH) technology to analyse genetic abnormalities in day-3 blastomeres and confirming it in trophectoderm biopsy showed concordance between day 3 diagnosis and day 5 reanalysis; Treff and coauthors showed more reliable results for SNP-microarray (96% vs. 83%) and also a lower mosaicism degree (31%) for SNP-microarray samples in a study comparing array technology versus FISH technique (18). These data would support the suggestion of some authors, who proposed that the incidence of mosaicism may have been overestimated in previous studies due to technical inconsistency of the FISH technique (17, 18, 19). At present, this matter remains controversial. Regarding pregnancy rates, in both types of biopsies higher pregnancy rates are obtained comparing with the control group, in which no biopsy was performed (14, 19). To sum up: DAY 5. TROPHECTODERM BIOPSY The blastocyst stage is currently supposed to be an optimal time to perform biopsies for PGD/PGS. The combination of improved blastocyst culture, trophectoderm (TE) biopsy, refined cryopreservation techniques, and molecular assays, such as array comparative genomic hybridization that allows for 24-chromosome screening, have led to a renaissance of PGS. TE biopsy will not detect every circumstance in which the embryo is at risk of aneuploidy, but it will detect mosaicism more reliably than cleavage-stage biopsy (which cannot be relied on at all for this purpose) (20, 21). Moreover, when diagnosing monogenic disorders in single blastomere cells using PCR-based protocols, there is a high risk of PCR failure due to either no amplification (allele dropout) or preferential amplification of one of the alleles, potentially resulting in a reduced number of unaffected embryos available for transfer. Increasing the amount of starting DNA template should in principle increase the sensitivity and reliability of genetic diagnosis. Therefore, the biopsy of multiple trophectoderm cells from the blastocyst rather than a single cell from cleavage stage embryos should potentially lead to improved PGD outcome for patients (14). How many cells should be removed? Research to determine the appropriate number of biopsied TE cells in blastocyst biopsies are limited. The exact number of biopsied TE cells is hard to count visually because cells are small and usually remain as a clump. In most studies using comparative genome hybridization or single-nucleotide polymorphism array technology for genetic testing, biopsied TE cells were used for genome amplification and their number was impossible to know. Moreover, some studies showed that removing four to five cells leads to better results. Therefore, the biopsied cell number should be higher in the blastocysts with better TE quality than those with worse characteristics (22, 23). Can biopsies affect blastocyst development and its implantation? Whereas it remains possible that biopsy of cleavage-stage embryos can critically arrest further development through reduction of cell mass, the low miscarriage rates and high term birth rates in the present series, as well as data presently under analysis, suggest that this is not the case for TE biopsy. It can be speculated that the damage to blastocyst development potential caused by TE biopsy would be less for blastocysts with a greater number of TE cells (21, 22). Some experts assured that TE biopsy at the blastocyst stage had no meaningful impact on the developmental competence of the embryo as measured by implantation and delivery rates. This contrasts with the information above-mentioned on the significant reduction in the probability for an embryo to implant and progress up to delivery (16). When combined with TE biopsy and blastocyst vitrification, SNP microarray has resulted in high implantation and low miscarriage rates for some IVF patients (15, 16, 24). Are there any limitations? Owing to the limitations of genetic analysis, most of the biopsied blastocysts need to be cryopreserved by vitrification, and blastocysts with normal results would be transferred in the next frozen cycle. In addition, biopsy of numerous cells from blastocysts with grade B or C may cause damage to the embryo, leading to either its arrest or implantation failure. However, 1-5 cells may be the appropriate biopsied TE cell number to maintain the implantation potential (15, 22). Also, the personnel experience of different embryologists is an influencing factor in this technique. The number of biopsied cells in the blastocyst biopsy is hard to quantify and largely dependent on the experience of embryologist (22). To sum up: WHAT CAN WE CONCLUDE? The availability of new embryology and molecular techniques allow preimplantation genetic diagnosis laboratories to offer patients at genetic risk the transfer of developmentally competent embryos, unaffected by genetic disease. Cleavage stage biopsy allows for fresh embryo transfer after genetic diagnosis. However, there are reports of high levels of mosaicism when the biopsy is performed on day 3. Trophectoderm biopsy, in turn, provides sufficient material for an effective and more reliable diagnosis in embryos compared to those on cleavage stage. Moreover, it seems that it does not compromise embryo implantation and pregnancy rates in PGD cycles. The drawback for this option is the usual need for cryopreservation and transfer in a different cycle. The offer of PGD in fertility centres has increased over the last decade, primarily due to the progress on the application of diagnostic methods. The choice for either development stage relates to successful outcomes in the clinic, which mainly depend on technical challenges and timing of the developing embryo. For the embryologists, both day-3 and day-5 approaches are supported by evidence, but it will be essential to consider every single aspect of them to evaluate the best option for the laboratory. REFERENCES:
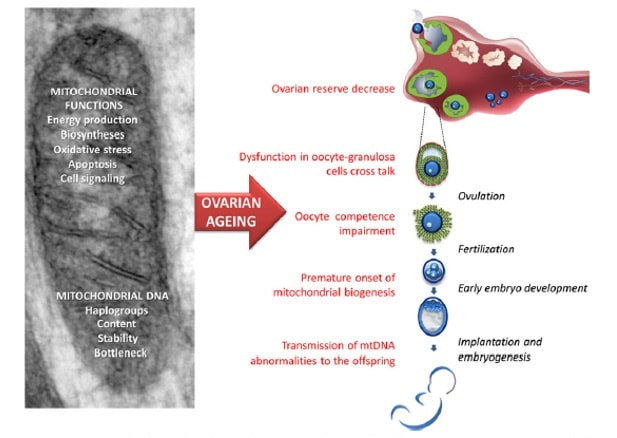
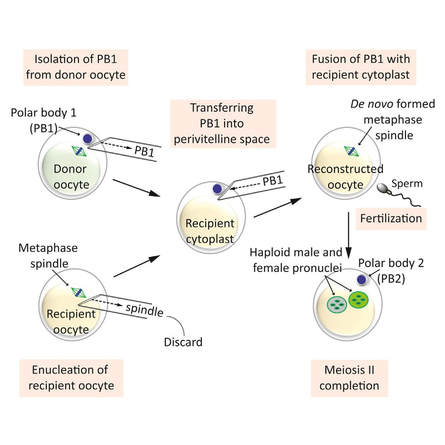
Authors: Shuyana Deba, Javier Del Río and Sara Sanz Special collaboration: Álvaro Martínez Moro Infertility affects millions of couples all around the world. In spite of the solutions to their problems reproductive technology can achieve, the efficacy is eventually limited by the number and the quality of the oocytes available from the woman. In actuality, such efficiency is determined by the ovarian reserve, the oocyte quality and the maternal age, among the most important factors (2). Diminished ovarian reserve (DOR) Since ovarian reserve defines the quantity and quality of the primordial follicle pool, diminished ovarian reserve (DOR) indicates a reduction in quantity in women of reproductive age. Consequently, it represents important cause of infertility in many couples. Moreover, DOR may be associated with low pregnancy rates and high pregnancy loss regardless of age, but further research is needed in order to fully understand its implications (3). Advanced maternal age It is well known that women’s fertility declines sharply after age 35 due to several factors, which include specific issues of reproductive organs (uterus and oviducts), general health and decreasing number and quality of oocytes over time. The oocyte pool starts to decline during foetal life and continues within the reproductive life of women. Oocyte quality also decreases as a consequence of the increased rate of aneuploidies observed with age: 74% at the age of 41–42, and up to 93% after the age of 42 (5). Advanced age is too associated with a reduction in the quality of the oocyte cytoplasm (ooplasm), which directly affects oocyte maturation (3). What are the main reasons for this reduction in ooplasm quality? Mitochondria are one of the most important organelles, which are affected in different ways (6,7): - Morphological and functional abnormalities - Mitochondrial swelling - Alterations in mitochondria's cristae - Vacuolization - Alterations of the membrane potential - Alterations of the metabolic pathways in cummulus cells, which may result in impaired mitochondria biogenesis during oogenesis. These effects are due to the higher ratio of mutation consequence of the proximity of these organelles to the respiratory chain, the inefficient repair mechanism and the exposure of histories. How these changes affect oocyte quality (8)? First of all, negative effects on chromosome segregation have been observed as a result of a decreasing ATP concentration (9,10). Additionally, defects have been found in different signalling pathways such as Ca2+ signalling, which affects fertilization and the subsequent embryo development (11). Nevertheless, different mitochondrial haplogroups should be taken into consideration. These have different bioenergetic functions, including production of reactive oxygen species (ROS) and mitochondrial coupling efficiency, aspects that might affect the oocyte longevity (13). Consequently, new techniques are being developed in order to increase the reproductive options in women with oocyte problems. Recently, one of these techniques that have been highly treated in the media is the development of additional viable oocytes from polar body genomes (2). HOW DOES TRANSFER OF POLAR BODY GENOME WORK? Originally, the transfer of polar body has been applied to cases of infertility with a genetic cause, such as the presence of mitochondrial diseases. These cases can be treated with the use of donor oocytes in clinical practice. Additionally, another application is the formation of human metaphase II (MII) oocytes, which increases the number of available oocytes for an assisted reproduction cycle (2). Two specific combined steps are needed. First, the donor oocyte spindle is removed, which requires the utilization of polarized light. Once located, it will be biopsied, obtaining an enucleated oocyte (14,15,16). Secondly, the patient polar body is biopsied, provided elimination of the spindle apparatus has been confirmed. Once both processes have been performed, the last step is the introduction of the polar body genome inside the enucleated oocyte (17). FUNCTIONAL HUMAN OOCYTES GENERATED BY TRANSFER OF POLAR BODY GENOMES Hong Ma and his group have tried to test the efficiency and possible limitations of this technique (ref). The main objective to be achieved was the formation of spindles resembling those typical of MII oocytes, including the appropriate chromosome dosage. HOW EFFICIENT IS THIS TECHNIQUE? Although DAPI staining demonstrated that all polar body nuclear transfer (PBNT)-oocytes contained spindle-chromosome complexes, only two of five experimental oocytes formed metaphase spindles similar to intact MII oocytes. This low number may be due to residual meiotic activity in enucleated human MII oocytes, which is sometimes not enough to induce formation of normal MII-like spindles. For a different cohort of oocytes, the rate of successful fertilization was 76%, still slightly lower than control oocytes. Furthermore, 42% of embryos reached blastocyst stage, indicating that most of the PBNT-oocytes were capable of completing the second meiotic division. Short tandem repeat (STR) analysis revealed that two sampled PBNT-blastocysts contained normal diploid chromosomes, determining that these embryos were completely viable. WHAT CAN BE CONCLUDED? • Polar body genome transfer seems to be a significant technique for the improvement of assisted reproductive technology (ART) outcomes and pregnancy rates, particularly for women with decreased ovarian reserve and low response to stimulation. • The cytoplasm from young donor oocytes may reduce incidences of low cytoplasmic oocyte quality. • It could provide an additional technique to support mitochondrial replacement therapy. Nevertheless, this technique is not suitable for women who cannot produce mature oocytes, typical profile of ART patients. Additionally, incidences of aneuploidy resulting from errors in mitosis or in the second meiotic division may still occur because of women advanced age. Larger datasets from this technique are needed to confirm its efficacy and safety. Also, improving preimplantation genetic screening (PGS) is critical before eventual clinical application. REFERENCES:
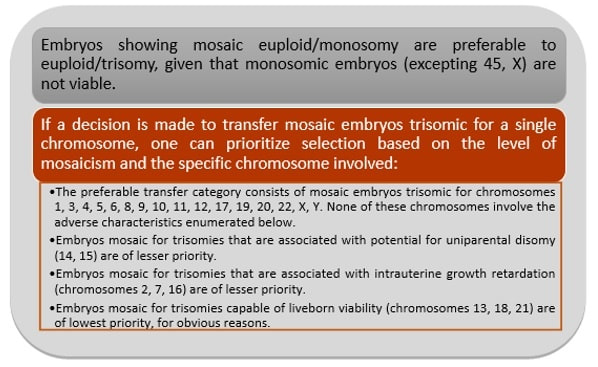
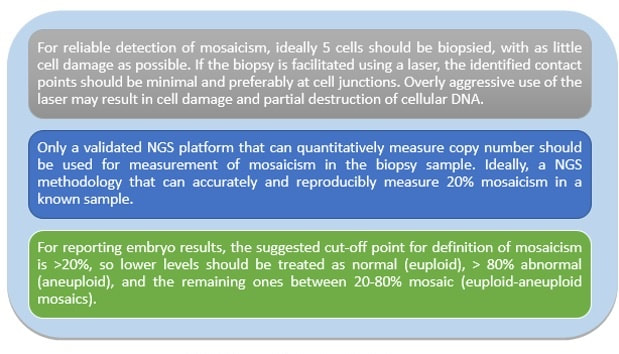
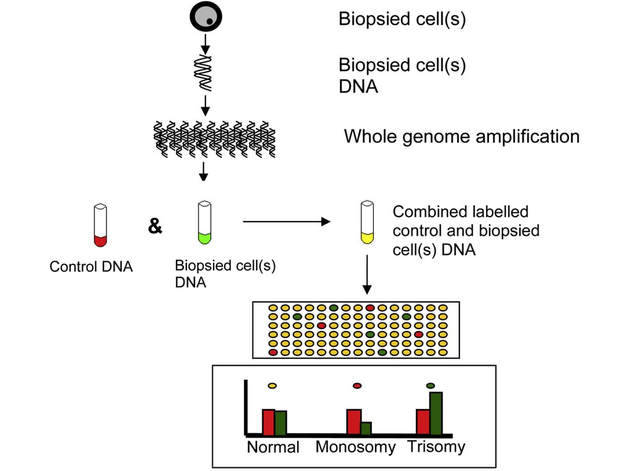
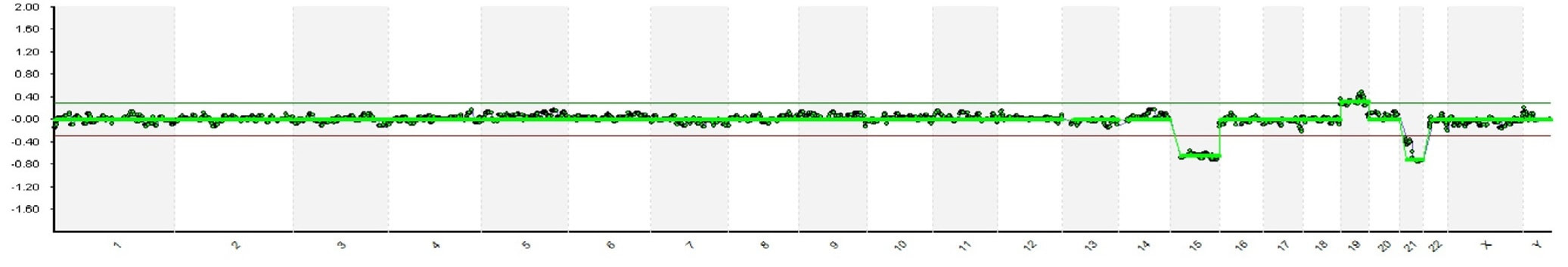
Authors: Javier Del Río and Sara Sanz Genetic problems in the embryo are one of the most important causes of pregnancy loss and miscarriage. However, identifying embryo mosaicism as the cause of genetic problems during development is not an easy task. WHAT IS A MOSAIC EMBRYO? The term mosaicism refers to the presence of more than just one cell line, which present different chromosome count (1). Additionally, the most common situation in these cases is the presence of a mixture of distinct aneuploid cells, rather than of a variety between euploid and aneuploid cells. These are the embryos that may be at risk of misdiagnosis (2). There are four possible types of mosaic embryos (3,4): 1. Embryos with a mix of euploid and aneuploid cells in the trophectoderm (TE), and with aneuploid cells in the inner cell mass (ICM). 2. Embryos with a mix of euploid and aneuploid TE cells and euploid ICM. 3. Embryos with euploid TE cells and aneuploid ICM. 4. Embryos with aneuploid TE cells and euploid ICM. Even though there exists no specific cut-off to determine mosaicism, the Preimplantational Genetic Diagnosis International Society (PGDIS) suggests an embryo with more than 20% of aneuploid cells to be considered as mosaic. This means lower levels of mosaicism should be treated as normal (euploid) (5). It has been traditionally thought that only genetic problems in the oocyte or the sperm could be responsible for embryo mosaicism. Nevertheless, it is currently postulated that this also occurs during the first mitotic divisions, when maternal transcripts control the cell cycle of the early embryo (6). NORMAL INDICATIONS FOR CHROMOSOMAL TESTING TO DETECT EMBRYO MOSAICISM 1. Advanced maternal age. This is the most common cause for aneuploid problems (7). A recent study has shown that women between the ages of 35 and 43 years have more probabilities (an increase ranging from 28 to 78%) of presenting mis-segregation for the most clinically relevant aneuploidies, namely chromosomes 13,16,18,21 and 22 (8). 2. Severe male factor infertility. Even though levels of sperm aneuploidy are associated with increased levels of chromosomal abnormalities in embryos (9), such abnormalities could also arise from certain males who do not present any chromosomal abnormality a priori. Such could be specific cases of oligoastenozoospermic patients (10). 3. Recurrent implantation failure (RIF). In spite of the lack of specifications for such diagnosis, it is usually defined as the occurrence of three or more failed IVF attempts due to an unidentified cause. RIF is the usual diagnosis in those cases in which after a cumulative transfer of more than 10 good-quality embryos, the eventual result is IVF failure (7,11,12,13). 4. Recurrent miscarriage. The definition of this concept may vary for every country. However, generally speaking it can be defined as the occurrence of 3 or more consecutive miscarriages once pregnancy has reached at least 14 weeks (14). The main cause for this problem seems to be aneuploidy, which has been identified as the leading cause in a high percentage of miscarriages (15,16). 5. Previous trisomic pregnancy. Cases in which there has been a previous trisomic pregnancy entail higher probability of suffering from another aneuploid conception. Therefore, it is in this group of patients in which it would be beneficial to conduct a study to find out possible related causes (17). BIOPSY TECHNIQUES TO STUDY A CASE OF MOSAICISM Although there are different biopsy strategies, depending on the embryo stage, blastocyst biopsy is recommended over both polar body and blastomere biopsies in those cases in which mosaicism is suspected. Blastocyst biopsy is less invasive, it is possible to extract a higher number of cells, which increases the probability to confirm mosaicism, and it is cheaper than the other techniques due to the lower number of embryos required for biopsy (7). Furthermore, if cells with different chromosome complements are widely distributed throughout the trophectoderm, there might be a good chance of capturing a representative sample. If cells are clustered, mosaicism could not be easily detected, thus providing false normal results (18). TECHNIQUES EMPLOYED TO DIAGNOSE MOSAICISM The earliest trials of PGD involved the use of karyotyping and PCR. By the mid-90s, the use of cytogenetic techniques such as FISH allowed for the progress of preimplantation diagnosis. This very own approach was later shown to impose important technical limitations to the analysis, and so it was encouraged the development of new technologies that could minimize the errors in diagnosis (19). Fluorescence in situ hybridization (FISH) It allows for the analysis and identification of chromosomes or chromosome fragments with 5-10 fluorescently labelled molecular probes from one cell (blastomere) (Fig. 2). This cell can be biopsied from day-3 embryos, it could belong to the trophectoderm from blastocysts or it could also be a polar body biopsied from an oocyte or a zygote. Therefore, PGS–FISH diagnosis is limited to the most common abnormalities involving chromosomes 13, 15–18, 21, 22, X and Y. Some studies of preimplantation embryos diagnosed by using this technique estimate a 5-7% error caused by mosaicism when embryos are reanalyzed. Additionally, FISH is being rapidly replaced by other DNA analysis methods with higher efficiency (20,21). Array Comparative Genomic Hybridization (aCGH) This technique relies on whole genome amplification from one or more blastomeres (Fig. 3). It provides a quantitative analysis based on the comparison between the relative amount of tested DNA and the control DNA. Thus, chromosome imbalances such as aneuploidies, unbalanced translocations, deletions and duplications are easily detected. However, since balanced chromosome rearrangements such as reciprocal translocations or inversions do not affect copy number, such alterations cannot be identified (19,20). When blastomeres are analyzed by aCGH, the error rate measured is merely 2% (21,23). Notably, some studies have found that PGS-aCGH after blastocyst biopsy provides higher implantation and pregnancy rates than PGS-FISH (24). Next Generation Sequencing (NGS) Next Generation Sequencing belongs to the group of Massively Parallel Sequencing (MPS) methods that allow for parallel processing of an extremely large number of nucleic acid molecules (Fig. 4). As a result of sequencing on a microspace scale, it has been possible to drastically increase the amount of information collected during one test up to an entire human genome. Also, it is the only method that allows for analysis of all chromosomes (aneuploidies or translocations) and mutations responsible for any single-gene disease, just using one biopsy and in a single step (20). Although clinical results have documented high pregnancy rates following transfer of screened embryos, further data along with an extended use in clinical application are required to better define the role of NGS in PGS. Nevertheless, it seems that this method may actually lead to reduced costs per patient, thus allowing IVF couples a wider use of PGS for choosing the most competent embryo for transfer (26). DIFFICULTIES WHEN MANAGING RESULTS The detection of mosaicism at an early stage does not mean that it will spread along embryo development (27). However, the utilization of chromosome identification techniques as part of the IVF process makes it possible to identify embryos “at risk of mosaicism’’ in order to select those that are suitable for transfer (18). Mosaic embryos are supposedly less competent than others due to a reduced implantation potential. Therefore, by discarding mosaic embryos implantation rates should be improved and, simultaneously, embryo loss rates reduced. Nevertheless, mosaic embryos may still have reproductive potential, and consequently they could still be viable. Furthermore, discarding embryos capable of producing healthy children will decrease pregnancy rates in those patients who get a low number of blastocysts in the pool of transferable embryos (18). It is important to take into account the reaction of the patients when they are informed about their embryos being at risk for mosaicism, what may entail genetic abnormalities, reduce implantation rates, increase loss risk and even diminish obstetrical and neonatal outcomes. However, there is not a simple answer when patients decide to transfer a mosaic embryo; either way, the obstetrical team should be informed for future screens (18). SUGGESTED GUIDELINES BY THE PREIMPLANTATINO GENETIC DIAGNOSIS INTERNATIONAL SOCIETY (PGDIS)
This article has been selected for publication in the Scientists in Reproductive Technologies (SIRT) Newsletter of The Fertility Society of Australia: DEL RÍO, J. and SANZ, S. (2017) Mosaic embryos are capable of producing healthy children. How to handle it? Fertility Society of Australia - SIRT Newsletter 4(20): 12-15. REFERENCES
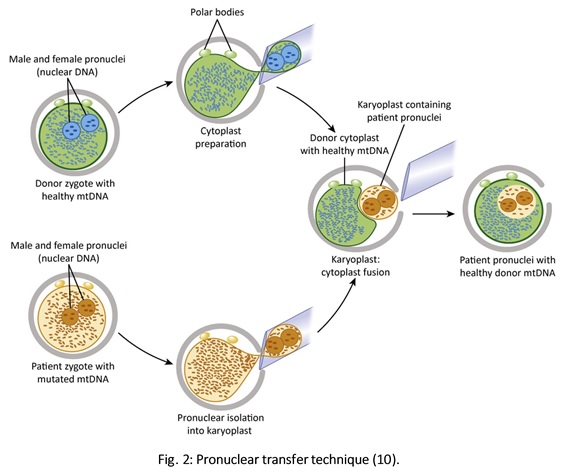
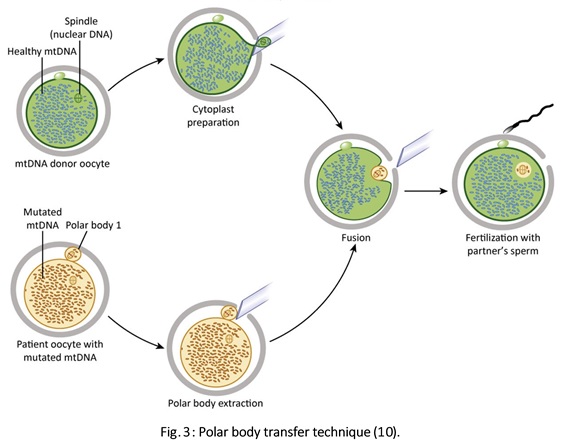
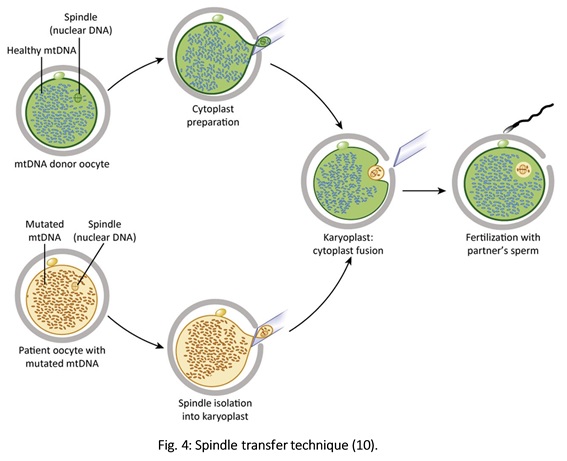
Authors: Javier Del Río and Sara Sanz Although most of the genetic material of eukaryotic cells is located inside the nucleus, mitochondria are organelles that also possess a certain amount of DNA. Mutations in mitochondrial DNA (mtDNA) or nuclear genes involved in mitochondrial function are cause of infertility and diseases, not only in individuals, but also in their offspring. In these cases, one of the solutions known to be efficient in order to conceive and give birth to a healthy child is the "three-parent in vitro fertilization" approach. THE MITOCHONDRIAL GENOME IS MATERNALLY INHERITED During fertilization, mitochondria from sperm are normally eliminated by a ubiquitin-dependent mechanism. As a consequence, in case the father carries the mutation both his health and fertility could become affected, but never his offspring (2). By contrast, mitochondria in the oocyte must present a specific location and distribution pattern (which represents an actual sign of oocyte maturation), and they are solely inherited from the mother. WHY ARE MUTATIONS IN mtDNA OR IN NUCLEAR GENES INVOLVED IN MITOCHONDRIAL FUNCTIONS SO PROBLEMATIC? Mitochondria provide energy to the cells through oxidative phosphorylation, and so mutations in their genome mainly affect structures form the nervous system, heart, skeletal muscle, pancreas, gonads, colon, blood, kidney or liver (3). Why these structures? The higher the energy demand is, the higher the need for more mitochondria in the cells (4). Also, cells that present a slower division process are more likely to present some kind of mtDNA mutation (5). HOLOPLASMY vs. HETEROPLASMY The situation in which all cells from an individual contain identical mtDNA (mutated or otherwise) is known as holoplasmy. By contrast, heteroplasmy is defined as the condition in which part of the mitochondria from the same individual present a DNA content that is different from the other. These cases are the most common among patients affected by mitochondrial DNA diseases (6). WHY DOES HETEROPLASTY REPRESENT A PROBLEM? Even though heteroplasmy implies the presence of two different DNA contents in the cell, cells with great amounts of mutant (or affected by a specific condition) mitochondria respond to proliferate their entire DNA. This is why the percentage of mutant/affected mtDNA tend to increase in certain tissues (7). THE "BOTTLENECK EFFECT" AND THE "THRESHOLD EFFECT" During oogenesis, only a subset of molecules of mtDNA are eventually amplified and passed on to the offspring (8). This effect explains why it is possible to obtain homoplasmic individuals in just a few generations (2). Previous reports on human diseases caused by an mtDNA mutation have shown that the mutation needs to be present at a certain percentage in order to manifest pathological effects. Typically, this percentage should be higher than 60-80% (8,9), although it also depends on age, affected tissues, type of mutations, etc. (4) WHY TO APPLY THE "THREE-PARENT IN VITRO FERTILIZATION" APPROACH? It might be reasonable to think of other possibilities to treat patients suffering from mitochondrial diseases in order to achieve pregnancy. Rather than prenatal diagnosis or preimplantation genetic diagnosis, the "three-parent in vitro fertilization" technique because (4): In the first case: 1. It needs a uniform mtDNA distribution in the extra-embryonic and foetal tissues. 2. It needs the mutant DNA load to remain constant over time. 3. There must be a close relation between the severity of the disease and the amount of mutant DNA. As for preimplantation genetic diagnosis (6,9): 1. It is not applicable to patients with high levels of heteroplasmy. 2. It reduces but does not eliminate the risk of suffering from a mitochondrial disease-related condition. 3. The amount of tDNA found in blastomeres or the trophectoderm does not represent the whole embryo. 4. This approach is not an efficient diagnosis due to the combination of heteroplasmy and the "bottleneck effect". KNOWB TECHNIQUES TO BE USED FOR MITOCHONDRIAL REPLACEMENT Pronuclear transfer It involves the transfer of the two pronuclei from a zygote affected by diseased (or mutated) mitochondria into an enucleated zygote containing healthy mitochondria. Even though this technique has not yet been performed in humans, the efficiency of pronuclear transfer in mice has been adversely affected by descendants bearing high levels of carryover mtDNA (10). Polar body transfer Since the polar body has a lower proportion of mitochondria around it, this is currently considered the best method for preventing the transmission of mutated mtDNA on to the next generation (9,10). Embryos derived from polar body transfer support normal fertilization and are capable of producing live offspring in mouse. Polar body transfers leading to a minimal amount of affected mtDNA carryover have demonstrated the great potential of this technique for preventing inherited mitochondrial DNA diseases (9,10). When applied in mice, this technique has shown the best success rate so far due to the transfer of mitochondria being lowered to a minimum (9). Spindle transfer This technique involves transferring the meiotic spindle along with the associated chromosomes, the spindle-chromosome complex (SCC), from an unfertilized oocyte with affected mitochondria into an enucleated healthy mitochondria-containing oocyte (11). This technique recently became popular when performed by Dr. John Zhang and his team, hitting the media within the latest weeks. However, potential problems could arise; just as for the previous techniques, the spindle is also surrounded by mitochondria, and so they could too be introduced into the ooplasm, thus causing heteroplasmy (12). WHAT DOES LAW STATE REGARDING MITOCHONDRIAL REPLACEMENT? So far, scientific societies are very skeptical about experimental techniques. Thus, this particular approach is specifically prohibited by the Food and Drug Administration (FDA) in the US. It can only be performed in countries such as Mexico, where legislation is more flexible, or in UK, where it was approved for application in very specific cases (14). CURRENT DATA ON MITOCHONDRIAL REPLACEMENT As it has been previously mentioned, cases of cytoplasm transfer have been performed. In fact, there have been around 30-50 live births from this technique. However, newborns presented certain genetic defects, and so this technique was banned and replaced by pronuclear transfer and meiotic spindle transfer (14). Such data demonstrate the potential damage that could be inflicted to the embryo when performing these techniques (12,15). In addition to this, ethical issues must also be taken into account, which means the sole possibility of successfully applying a specific procedure does not imply its moral appropriateness. In order to guarantee so, a committee of experts should pronounce their opinions and reach a consensus about it. On a related note, long-term effects derived from these procedures are still unknown, and so it would be necessary to monitor all babies born through these techniques. This post has been published in the Scientists in Reproductive Technologies (SIRT) newsletter, a special interest group representing the scientific membership of The Fertility Society of Australia. REFERENCES
|
Entries
March 2019
Categories
All
 2016-2019. All Rights Reserved by Embryologist Media. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License . |
Embryologist Media